Simeon Rossmann
Forsker
Biografi
Jeg forsker på forskjellige anvendte aspekter innenfor plantepatologi. Jeg har jobbet mye med Pectobacteriaceae, som forårsaker bløtråte og stengelråte i potet og ulike andre sykdommer i mange andre vertsplanter. Jeg er spesielt interessert i insekter assosiert med disse bakteriene og andre plantepatogene bakterier i feltet og molekylære mekanismene bak disse forskjellige assosiasjonene.
I det siste var jeg fokusert på praktiske og bioinformatiske implementeringer av metabarkoding eller amplikonsekvensering som et verktøy for deteksjon og identifisering av plantepatogene sopp, Oomycota, nematoder, bakterier, og invaderende plante- og insektarter.
Jeg fikk BSc og MSc fra University of Tübingen, hvor jeg oppdaget min lidenskap for plantepatologi ved Center for Plant Molecular Biology (ZMBP). I løpet av doktorgradsperioden min (2015-2018) undersøkte jeg bløtråte i norske potet med prof. May Bente Brurberg (NIBIO / NMBU) som hovedveileder.
Sammendrag
Det er ikke registrert sammendrag
Sammendrag
Det er ikke registrert sammendrag
Forfattere
May Bente Brurberg Simeon Rossmann Erik Lysøe Monica Skogen Håvard Eikemo Paulina Paluchowska Mirella Ludwiczewska Sylwester Sobkowiak Marta Janiszewska Zhimin Yin Jadwiga SliwkaSammendrag
Det er ikke registrert sammendrag

Divisjon for bioteknologi og plantehelse
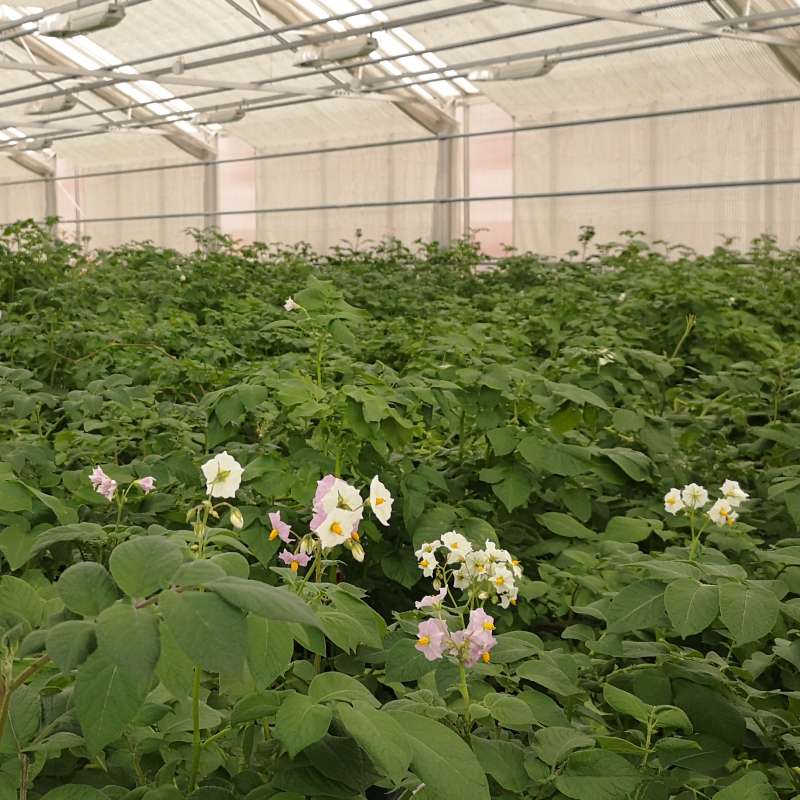
Genomisk kartlegging av sykdomsresistensgener i potet
Potettørråte er den mest ødeleggende sykdommen i norsk potetproduksjon og gir store avlings- og økonomiske tap. For å redusere pesticidbruk og sikre bærekraftig matproduksjon må nye sorter utvikles med varig genetisk resistens.
Divisjon for bioteknologi og plantehelse
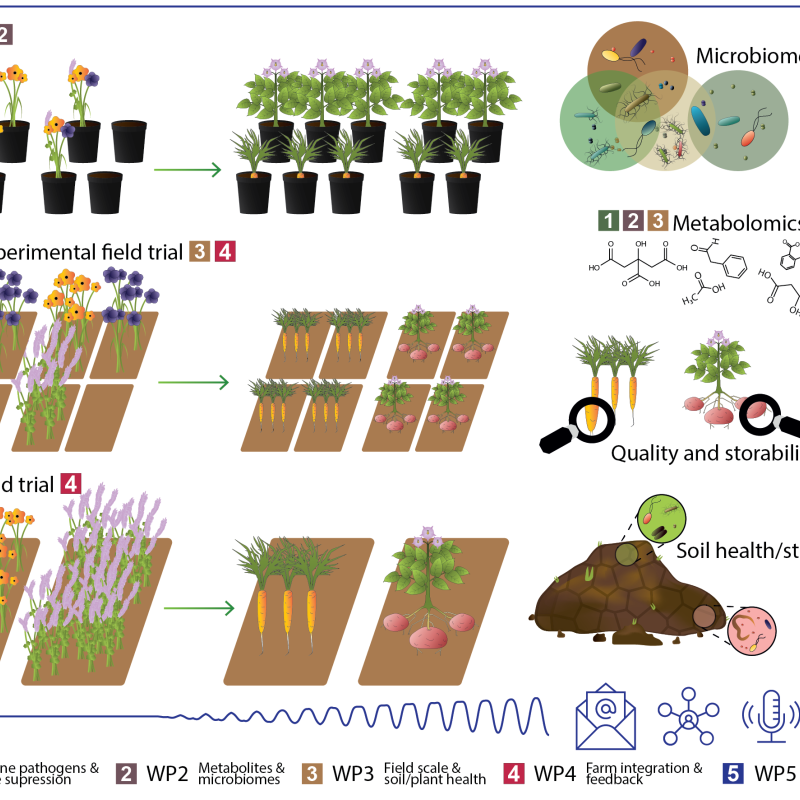
Cropdrive
Cropdrive aims to identify a selection of cover crops suitable for use in root vegetable and potato production with beneficial impacts on both soil and plant health, and greenhouse gas exchange.
